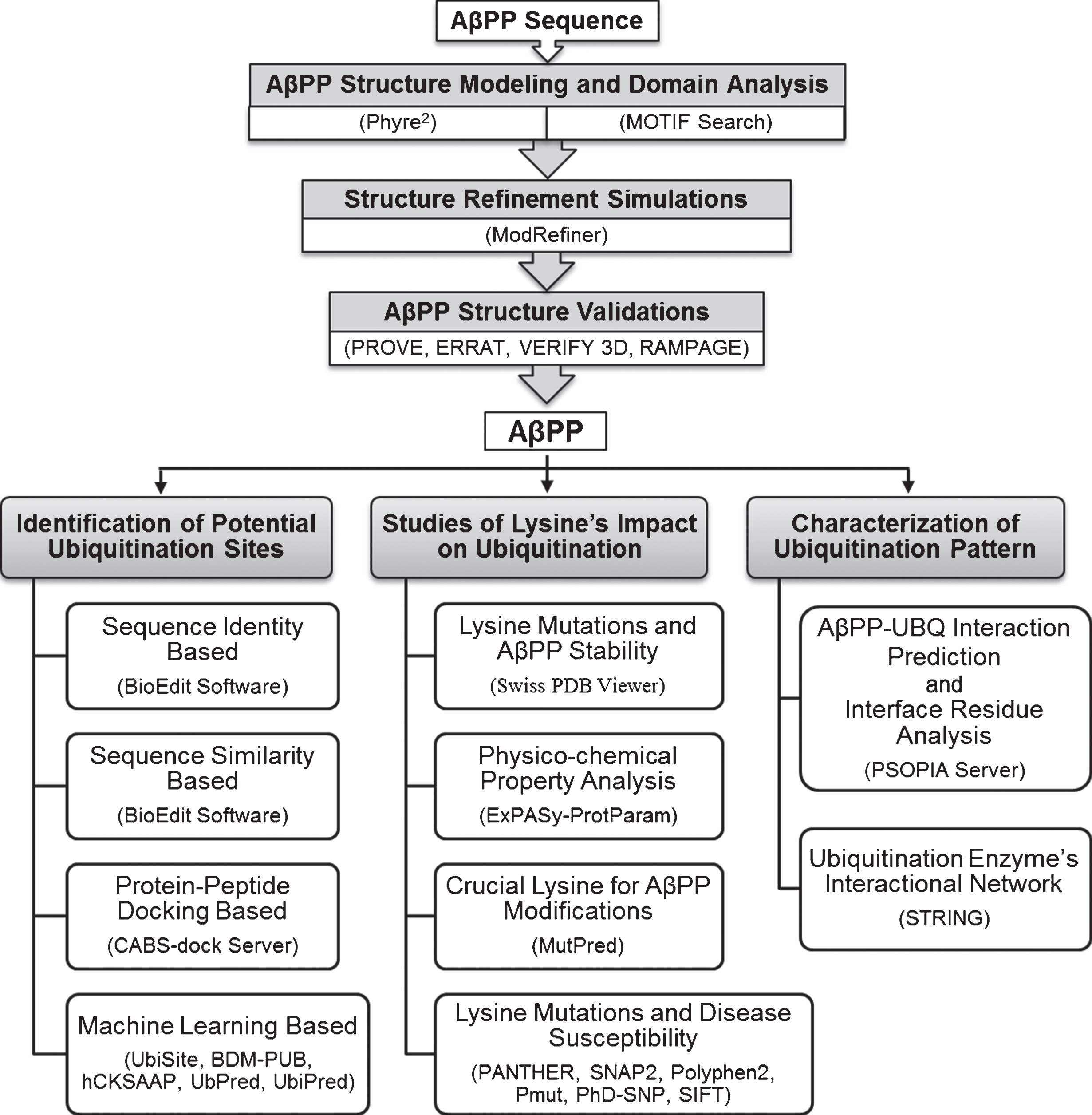
Here we provide a webserver and Python3 application that fixes the PDB sequence numbering problem by replacing the author numbering with numbering derived from the corresponding UniProt sequences. In addition to the coordinates, there are many fields that contain structural and functional information regarding specific residues numbered according to the author. For instance, some authors may include N-terminal signal peptides or the N-terminal methionine in the sequence numbering and others may not. This results in the same protein being numbered differently in the available PDB entries. However, these studies are made more difficult because authors are allowed by the PDB to number the amino acids in each protein sequence in any manner they wish.
Many proteins have been studied under different conditions, including binding partners such as ligands, nucleic acids, or other proteins mutations, and post-translational modifications, thus enabling extensive comparative structure-function studies. In mid 2021, the database has almost 180,000 structures solved by X-ray crystallography, nuclear magnetic resonance, cryo-electron microscopy, and other methods.

#INTRODUCTION TO BIOEDIT AND SWISS PDB VIEWER ARCHIVE#
The Protein Data Bank (PDB) was established at Brookhaven National Laboratories in 1971 as an archive for biological macromolecular crystal structures.
